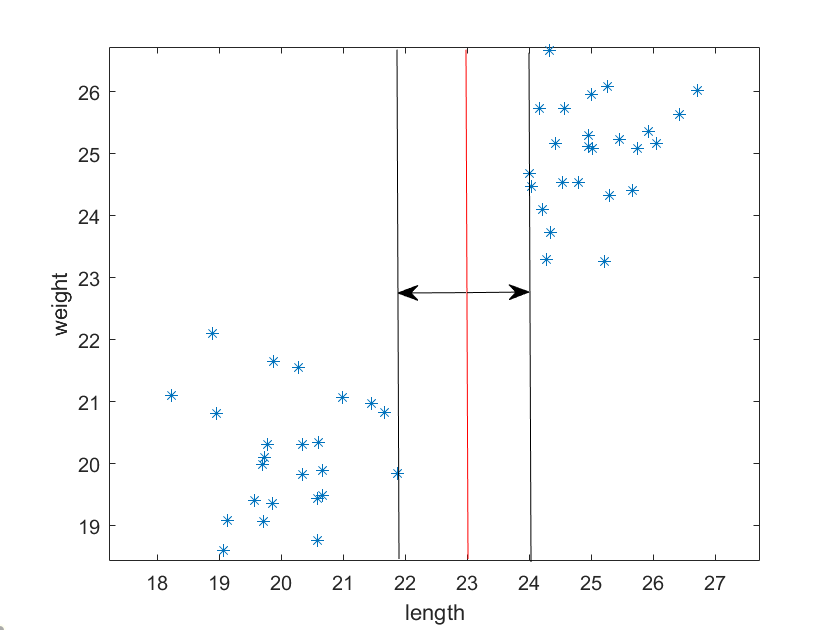
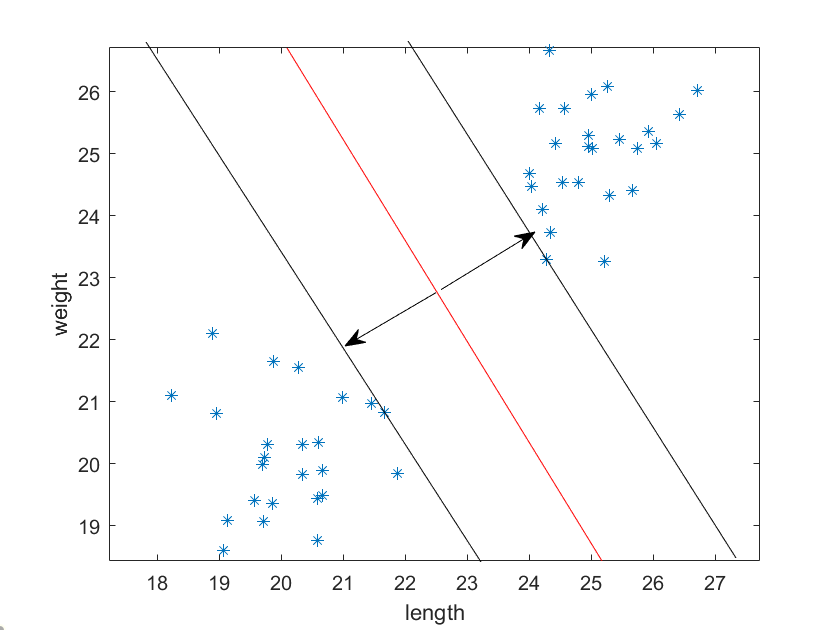
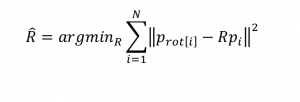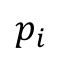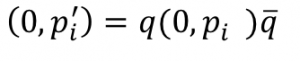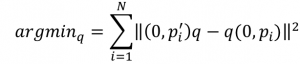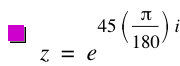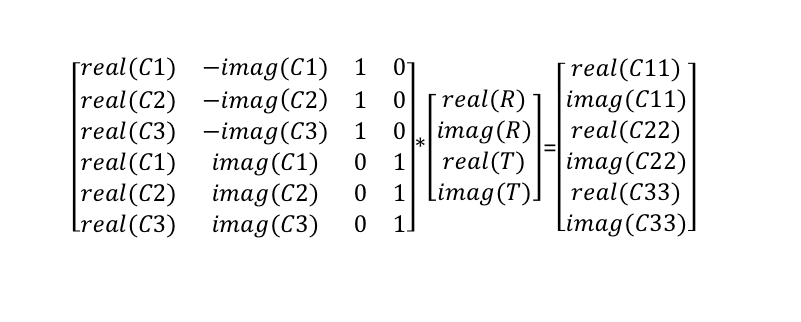In the previous post we discussed 2D – 2D image registration, and in this post we’ll continue that discussion with 3D – 3D image registration.
I’d like to again give credit to Dr. Joachim Hornegger’s online videos, available here, which led me to explore actually implementing these algorithms in Matlab.
In the case of 3D -3D Rigid Image registration, we have 2 sets of 3 points located in 3D space though prior measurements. In the case of surgery of the human body, we might have 3 small gold markers, known as “fiducial markers” in each 3d image. If our measurements of the marker locations were taken at different times, or with different types of imaging technology, we want to make sure that we line up the two 3d models as closely as possible, so we can accurately locate the features of interest.
Coordinate systems and rotation in 3D
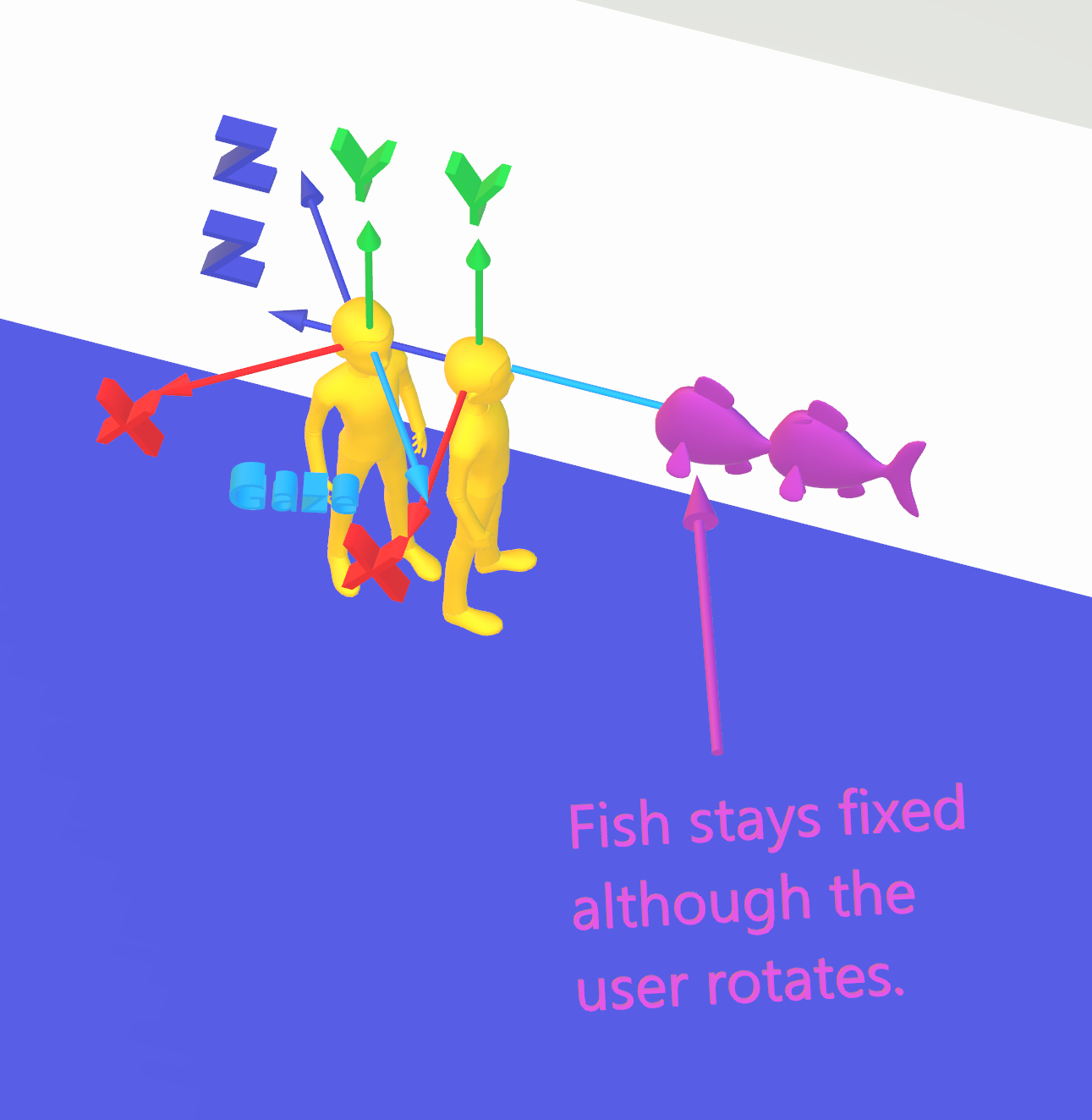
Most people are familiar with 3D coordinates as an extension of the 2D Cartesian coordinate system, by adding a Z axis to the X and Y axis. When rotating points in 3D, it is common and intuitive to think of rotating, for instance, 3 degrees about the X axis, -3 degrees about the Y axis, and 10 degrees about the Z axis.
However, there are limitations to this system that are severe enough to lead us to abandon the intuitive approach in favor of something more reliable. One limitation is that the order of the axes that you choose will change the final result – for instance, rotating first about X, then about Y, then about Z, will give a different final result than rotating first about Y, then X, then Z.
In addition, when you rotate at 90 degrees, the math to compute the final results can give ambiguous results.
This representation of rotations in 3D is often referred to as “Euler Angles”. I won’t go into the details of the limitations, but numerous examples can be found online that show the problems with using Euler Angles for computing complex rotational transformations.
One interesting related example happened on the Apollo 11 space craft:
http://en.wikipedia.org/wiki/Gimbal_lock#Gimbal_lock_on_Apollo_11
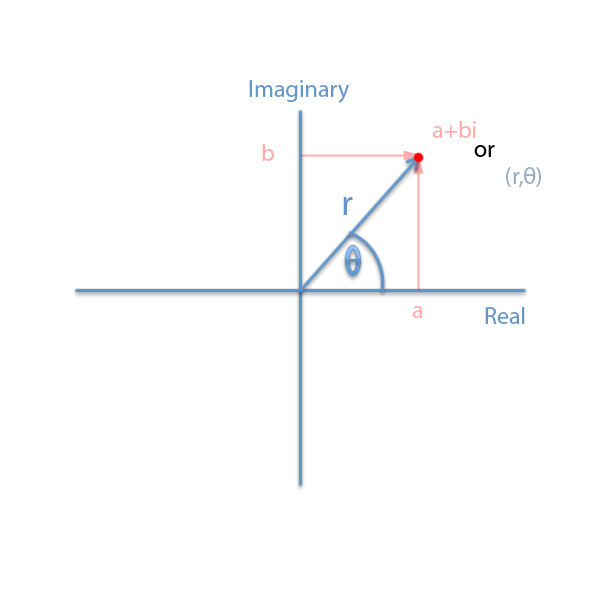
Quaternions
In my previous post we investigated using Complex numbers to make rotations simpler to solve using linear algebra. Quaternions provide us with a very similar advantage – they allow us to define rotations without the problems of Euler Angles mentioned above, and also provide us with a convenient structure to perform a least squares estimate of the rotation angle.
I won’t go into the details of Quaternions here, but you can find many articles and books that describe their use for defining rotations, especially in software like computer games.
For our purposes, I have written a Matlab class that encapsulates the operations of and on a Quaternion. The Quaternion consists of four member variables – w,x,y and z. If the w component is set to zero, we have a 3D vector represented within the quaternion – this makes it convenient to mulitply points times quaternions using the same object class.
The problem statement
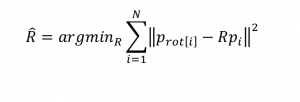
Source – Hornegger lecture notes, 2011, J. Hornegger, D. Paulus, M. Kowarschik
As stated pretty clearly above, the problem we are trying to solve is to find the rotation matrix R that minimizes the squared difference between the measured point
and the original point 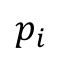 rotated by R.
rotated by R.
Since the rotation matrix R contains sin() and cos() functions, minimizing the equation is not a simple task, and would require an iterative, algorithmic solution. With Quaternions we can create a straightforward set of linear equations.
The rotation by quaternions can be represented as follows:
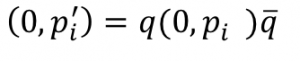
Recall that the left hand side, p’, is a quaternion with the w component set to 0. Likewise, on the right hand side we have the original point as quaternion p with it’s w component set to 0. The  character is the quaternion conjugate.
character is the quaternion conjugate.
We can factor out the conjugate by multiplying both sides by q:
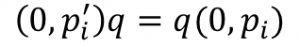
and the subtract the right side and we are left with
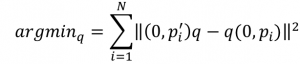
Source – Hornegger lecture notes, 2011, J. Hornegger, D. Paulus, M. Kowarschik
The above says we need to find the quaternion q that minimizes the sum of the squared differences between the rotated point (as measured) and the point to be rotated by q.
In other words, in a perfect world
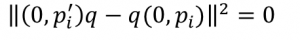
we would look for the quaternion q that makes the difference 0, or as close to it as possible.
In linear algebra terms, we’re looking for the nullspace of the matrix.
Setting up the matrix
I created a helper function in Matlab
MakeCoefficientMatrix( p, pPrime )
which returns a matrix made from p, the original 3D vector, and pPrime, the rotated vector.
We’re creating the matrix from
(ppPrime* q) – (q * p), so we need to define quaternion multiplication, which is as follows
quat.w = q1.w*q2.w – q1.x*q2.x – q1.y*q2.y – q1.z*q2.z;
quat.x = q1.w*q2.x + q1.x*q2.w + q1.y*q2.z – q1.z*q2.y;
quat.y = q1.w*q2.y + q1.y*q2.w + q1.z*q2.x – q12.x*q2.z;
quat.z = q1.w*q2.z + q1.z*q2.w + q1.x*q2.y – q1.y*q2.x;
So, given the above we can view the matrix as
(ppPrime* q) – (q * p) =
ppPrime.w*q.w – ppPrime.x*q.x – ppPrime.y*q.y – ppPrime.z*q.z – q.w*p.w + q.x*p.x + q.y*p.y + q.z*p.z;
We’re looking for q, so factor q out
q.w(pPrime.w – p.w) – qx(pPrime.x – p.x) – q.y(pPrime.y – p.y) – q.z(pPrime.z – p.z);
q.w(pPrime.x – p.x) + q.x(pPrime.w – p.w) – q.y(pPrime.z + p.z) + q.z(pPrime.y + p.y);
q.w(pPrime.y – p.y) + q.x(pPrime.z + p.z) + q.y (pPrime.w – p.w) – q.z(pPrime.x + p.x )
q.w(pPrime.z – p.z) – q.x(pPrime.y + p.y) + q.y(pPrime.x + q.y) + q.z(pPrime.w – p.w);
So the complete function is
function [ CM ] = MakeCoefficientMatrix( p, pPrime )
CM = zeros(4,4);
CM(1,1) = pPrime.w – p.w;
CM(1,2) = -(pPrime.x – p.x);
CM(1,3) = -(pPrime.y – p.y);
CM(1,4) = -(pPrime.z – p.z);
CM(2,1) = pPrime.x – p.x;
CM(2,2) = pPrime.w – p.w;
CM(2,3) = -(pPrime.z + p.z);
CM(2,4) = pPrime.y + p.y;
CM(3,1) = pPrime.y – p.y;
CM(3,2) = pPrime.z + p.z;
CM(3,3) = pPrime.w – p.w;
CM(3,4) = -(pPrime.x + p.x);
CM(4,1) = pPrime.z – p.z;
CM(4,2) = -(pPrime.y + p.y);
CM(4,3) = pPrime.x + p.x;
CM(4,4) = pPrime.w – p.w;
end;
Singular Value Decomposition to solve for nullspace
We can make the complete matrix from the 6 points as below, using the helper function:
CM = MakeCoefficientMatrix(p1,pPrime1);
CM2 = MakeCoefficientMatrix(p2,pPrime2);
CM3 = MakeCoefficientMatrix(p3,pPrime3);
% put together
CM = [CM;CM2;CM3];
[U S V] = svd(CM,’econ’);
The singular value decomposition of a matrix is probably one of the most important matrix factorizations to be familiar with. We can use this to find the null space of the matrix, and we can use it to solve other least squares types of matrix computations. The SVD will be a subject of a future blog post.
For now, we take advantage of the fact that the null space of the matrix is in the last column of V:
% get the null space
ns = V(:,4);
qanswer = Quaternion();
qanswer.w = ns(1,1);
qanswer.x = ns(2,1);
qanswer.y = ns(3,1);
qanswer.z = ns(4,1);
and we can check our results by plotting a cross against our rotated objects:
pPrime1 = qanswer * p1 * qanswer.Conjugate();
pPrime2 = qanswer * p2 * qanswer.Conjugate();
plt.PlotVector3WithMarker(pPrime1,pPrime1,’+’,[1 0 0],80,[1 0 0]);
plt.PlotVector3WithMarker(pPrime2,pPrime2,’+’,[1 0 0],80,[1 0 0]);
plt.PlotVector3WithMarker(pPrime3,pPrime3,’+’,[1 0 0],80,[1 0 0]);
I’ve created a movie showing 3 original spheres, which are then rotated by a random amount (which we don’t “know”),
and then using our method described above we find the angle of rotation and plot a cross hair on our “found” spheres.
3d rotation movie
The complete matlab code is below – note, I use my own object-oriented classes, so this code is only a model for your own code, but should be a good example to build from.
Rick Frank
clc;
clear all;
close all;
plt = Plotting();
hold on;
grid on;
colordef black;
% size of our 3d space…
axisVector = [-1.2 1.2 -1.2 1.2 -1.2 1.2];
sphereSize = 0.1;
% Origin and axis for display.
Origin = Vector3(0,0,0);
vAxisX = Vector3(1,0,0);
vAxisY = Vector3(0,1,0);
vAxisZ = Vector3(0,0,1);
% these points are the first measured points.
v1 = Vector3(-0.9,0.2,0.4);
v2 = Vector3(0.8,.9,0.5);
v3 = Vector3(0.4 ,0.4 ,0.3);
red = [1 0 0];
blue = [0 0 1];
green = [0 1 0];
colorX = [1 1 0];
colorY = [0 1 1];
colorZ = [1 0 1];
hold on;
plt.PlotVector3(Origin,vAxisX,’-‘,red);
plt.PlotVector3(Origin,vAxisY,’-‘,blue);
plt.PlotVector3(Origin,vAxisZ,’-‘,green);
plt.PlotSphere(v1,sphereSize,colorX,1.0,axisVector);
plt.PlotSphere(v2,sphereSize,colorY,1.0,axisVector);
plt.PlotSphere(v3,sphereSize,colorZ,1.0,axisVector);
vAxisRotation = Vector3(1,1,1);
d = randi([30,60])
% rotate n degrees about axis….
for n = 1:0.5:d
qr = Quaternion.MakeUnitQuatFromAngleAxis(n,vAxisRotation);
hold off
grid on;
plt.PlotSphere(v1,sphereSize,colorX,1,axisVector);
hold on;
grid on;
plt.PlotSphere(v2,sphereSize,colorY,1,axisVector);
plt.PlotSphere(v3,sphereSize,colorZ,1,axisVector);
p1 = Quaternion.Make3DVecQuat(v1);
%show it
plt.PlotSphere(p1,sphereSize,colorX,1,axisVector);
% rotate it
pPrime1 = qr * p1 * qr.Conjugate();
plt.PlotSphere(pPrime1,sphereSize,colorX,0.2,axisVector);
% another point
p2 = Quaternion.Make3DVecQuat(v2);
plt.PlotSphere(p2,sphereSize,colorY,1,axisVector);
pPrime2 = qr * p2 * qr.Conjugate();
plt.PlotSphere(pPrime2,sphereSize,colorY,0.2,axisVector);
p3 = Quaternion.Make3DVecQuat(v3);
plt.PlotSphere(p3,sphereSize,colorZ,1,axisVector);
pPrime3 = qr * p3 * qr.Conjugate();
plt.PlotSphere(pPrime3,sphereSize,colorZ,0.2,axisVector);
plt.PlotSphere(pPrime1,sphereSize,colorX,0.2,axisVector);
plt.PlotSphere(pPrime2,sphereSize,colorY,0.2,axisVector);
plt.PlotSphere(pPrime3,sphereSize,colorZ,0.2,axisVector);
plt.PlotVector3(Origin,vAxisX,’-‘,red);
plt.PlotVector3(Origin,vAxisY,’-‘,blue);
plt.PlotVector3(Origin,vAxisZ,’-‘,green);
set(gca,’CameraPosition’,[3,-4,-1]);
pause(0.01);
end
%
%(pqPrime1 * q) – (q * p1) = 0;
%(pqPrime2 * q) – (q * p2) = 0;
%etc….
% Utility to create the matrix
%if we add error, then what is the result of the singular values?
ErrorFactor = 0;
pPrime1.AddRandomError(ErrorFactor);
pPrime2.AddRandomError(ErrorFactor);
pPrime3.AddRandomError(ErrorFactor);
CM = MakeCoefficientMatrix(p1,pPrime1);
CM2 = MakeCoefficientMatrix(p2,pPrime2);
CM3 = MakeCoefficientMatrix(p3,pPrime3);
% put together
CM = [CM;CM2;CM3];
[U S V] = svd(CM,’econ’);
lastSingularValues = S(:,end);
delta = max(abs(lastSingularValues) – 0);
if ((delta > 0.00001) && (delta < 0.01))
disp(‘questionable measurements – adjusting’);
S(:,end) = 0;
newCM = U * S * V’;
[U S V] = svd(newCM);
else if (delta > 2.0e-16)
disp(‘bad measurements – should not be used’);
end
% get the null space
ns = V(:,4);
qanswer = Quaternion();
qanswer.w = ns(1,1);
qanswer.x = ns(2,1);
qanswer.y = ns(3,1);
qanswer.z = ns(4,1);
pause(1.0);
pPrime1 = qanswer * p1 * qanswer.Conjugate();
pPrime2 = qanswer * p2 * qanswer.Conjugate();
plt.PlotVector3WithMarker(pPrime1,pPrime1,’+’,[1 0 0],80,[1 0 0]);
plt.PlotVector3WithMarker(pPrime2,pPrime2,’+’,[1 0 0],80,[1 0 0]);
plt.PlotVector3WithMarker(pPrime3,pPrime3,’+’,[1 0 0],80,[1 0 0]);